
[ad_1]
Single-cell technologies are used to analyze genomic data, allowing scientists to better study organizational mechanisms and heterogeneity at the cellular scale. They also generate large amounts of extensive data, bringing cell biology into a new era.
At the Curie Institute, tumor specimens circulate between floors. They are first collected from patients by doctors in the basement, then sent to the pathology department for study. The specimens are analyzed cell by cell using revolutionary “single-cell” technology, then sent to another floor for sequencing. Finally, bioinformatics experts clean up the data sets generated by the sequencers and apply statistical methods to study them. The goal is to understand why tumors appear and how they resist treatment. Within a few months, living specimens become bodies of data. “Single-cell approaches have accelerated the development of Biology 2.0, where our work is now based on multidisciplinary collaborations, from doctors to biologists, data analysts and statisticians. Only with the participation of all these experts can the whole system work properly”, Research Professor at the Institut Curie, French National Institute of Scientific Research, Epigenetic mechanisms Breast cancer.
A scientist uses a multichannel pipette to prepare samples for single-cell analysis.
Since 2017, Vallot and his team have been using and developing single-cell technology for cancer research. This technology is an innovation that uses a process called microfluidics: cells pass one by one through microchannels and are encapsulated in oil droplets containing reagents. While encapsulated, the genetic material in the cells is labeled with a genetic barcode that identifies the cell. This approach allows scientists to study at the scale of individual cells and try to understand the cellular heterogeneity that exists within a single tumor. “Before the advent of single-cell technologies, we performed experiments on populations of millions of cells and then looked at the average expression of the genes involved.” She explained.
However, epigenetics experts point out that the single-cell scale is nothing new in biology. “When studying proteins, we use microscopy methods. The innovation of single-cell technology is that it is systematic, because we can focus on many cells and obtain relatively detailed information,” She added: “This new technology holds a lot of promise for cancer research and medicine, and presents a major opportunity: creating precision medicine, where each patient receives treatment tailored to their individual needs.”
Multiple methods
The first scientific publication applying single-cell methods to study RNA was in 2009. The technology gained momentum in 2015 and, starting in 2017, most French biological laboratories studying cancer are using it. In western Francewhich was launched in 2018. No bigger than a mini dishwasher, it offers countless possibilities for the team led by Stéphane Minvielle, research professor at CNRS and co-leader of the Integrative Cancer Genomics (ICAGEN) team. For the past two decades, the team at the Nantes University Hospital has been studying multiple myeloma, a cancer that affects the bone marrow. Their work focuses on the activity of cells in terms of messenger RNA: this is the transcriptomics approach, or the study and analysis of the transcription of DNA into messenger RNA.

Institut Curie has established the first single-cell analysis platform in France dedicated to cancer.
The combination of single-cell technology and transcriptomics is called single-cell RNA-seq. “Thanks to this, it is possible to distinguish between cell subtypes in the immune system, whereas before you could only see one group of cells. As a result, we can now observe the complex reactions involved and how tumor cells modify their environment,” Minvielle explains. Single-cell techniques can also be combined with other experimental protocols. Vallot studies epigenetics, which is the study of cell activity through the regulation of gene expression. “We are applying single-cell methods to epigenetics, which is something that only a few labs in the world are doing,” She explains that the combination of these two techniques is called single-cell ChIP-seq.
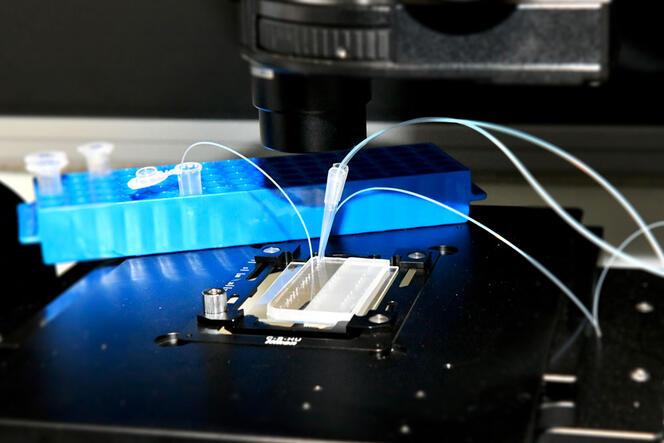

Institut Curie’s single-cell platform uses emerging droplet microfluidics technology to study cancer at the single-cell level. The platform allows the cells that make up a tumor to be individually analyzed to determine the right treatment for each patient.
exist IGFL of LyonIn southeastern France, Yad Ghavi-Helm is studying embryos of fruit flies, also known as DFruit fly. The goal is to use it as a model to understand the interactions between regulatory regions (enhancers) and genes in the genome. “Because mutations in enhancer sequences can affect the expression of the genes they regulate and lead to diseases such as cancer”” the scientist elaborates. As in Nantes, Yad Ghavi-Helm links single-cell RNA sequencing with spatial transcriptomics, a technology that combines high-throughput analysis of gene expression with information on the spatial localization of cells in their tissue or organ of origin.Until now, only imaging methods have allowed us to study gene expression in this way, but this technique has remained limited to a few genes in each experiment. With spatial transcriptomics, we can analyze hundreds or even thousands of genes at the same time,” said Dr. She added.
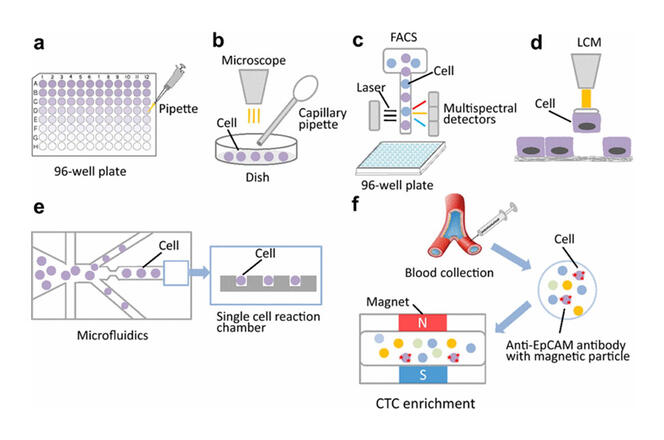
Different methods for isolating cells before analysis: a) Serial dilution. b) Mechanical manipulation. c) Laser microdissection. d) Flow cytometry. e) Isolating cells using microchannels connected to reaction chambers. f) Isolating cells using specific antibodies coupled to magnets.
Additionally, single-cell approaches allow for a deeper understanding of the role, modification, and degradation of proteins in cells. Proteomics is the science of studying the proteome, or all the proteins in a cell. Before the advent of single-cell technology, accessing these molecules was a major challenge: they are highly sensitive, cannot be amplified, and need to be analyzed on a very small scale, making traditional methods such as mass spectrometry both imprecise and inefficient. Starting at the single-cell scale, this technology applied to proteomics (SCP) can address questions that have not been explored by previous methods. Single-cell technology is developing so rapidly that its technical limitations are about to be overcome, but its cost remains a major obstacle for laboratories.
Turning life into an equation
“With the introduction of single-cell approaches, we have moved from Excel spreadsheets with 20,000 rows to computing servers,” Minvielle confirmed this. The technology has already expanded the scope of data generation: “Those who generate the information are no longer able to interpret it on their own, so biologists need support from engineers and researchers in statistical and artificial intelligence methods.”Statistics and machine learning are at the heart of the single-cell revolution, explains Franck Picard, a research professor at the Laboratory of Cell Biology and Modeling (LBMC) at CNRS and an expert in statistics and machine learning. Vallot couldn’t agree more:I feel like we’re only leveraging 5% of what this data is telling us. It would be great if we could analyze data as skillfully as ChatGPT does. But how can we equip ourselves with the resources and machines to get the most out of this data?” Single-cell approaches imply considerable interdisciplinarity, building bridges between biologists and those working in data analysis; thus attracting mathematicians into the field of biology is becoming a necessity.
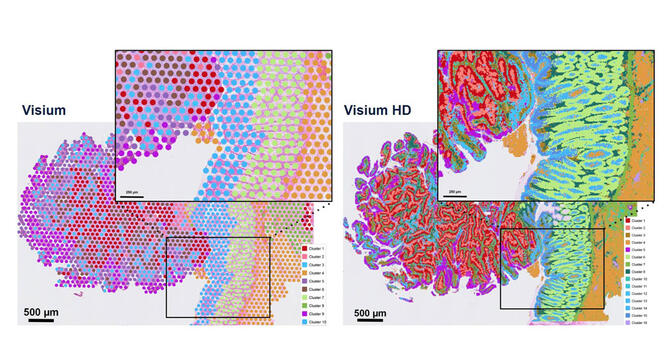
Gene expression profile of colorectal cancer tumors obtained by spatial transcriptomics. Each color corresponds to the expression of a gene of interest. The two graphs show the evolution of the resolution of the Visium technology developed by 10xGenomics. On the left: 55 µm resolution, on the right: 2 µm.
“The methodological challenge is to translate the organism into an equation.” Minvielle concluded. “Our advantage in France is that the training in mathematics and machine learning is very good.” Franck added. But competition in the field of artificial intelligence (AI) is fierce, as students are often recruited by the private sector, where they can earn much more than in public research. To address this problem, Picard and Vallot are looking to doctoral training. “We need to value and encourage PhD programs that foster these hybrid skills over the long term,” Picard confirmed that the goal is to develop more interdisciplinary institutes. In this dynamic, single-cell technology has been developed by German Institute of Health and Medical Sciences and InriaThe consortium brings together experts in single-cell data analysis. Piccard, who is coordinating a project of the initiative for the French National Center for Scientific Research, explained “Our goal is to develop new AI methods to exploit single-cell data. The long-term goal is to establish links with precision medicine and, above all, to ensure the emergence of a community specializing in this field in France”. The revolution has just begun. ♦
[ad_2]
Source link


